The goal of this notebook is to code a decision tree classifier that can be used with the following API:
df = pd.read_csv("data.csv")
train_df, test_df = train_test_split(df, test_size=0.2)
tree = decision_tree_algorithm(train_df)
accuracy = calculate_accuracy(test_df, tree)
The algorithm that is going to be implemented looks like this:
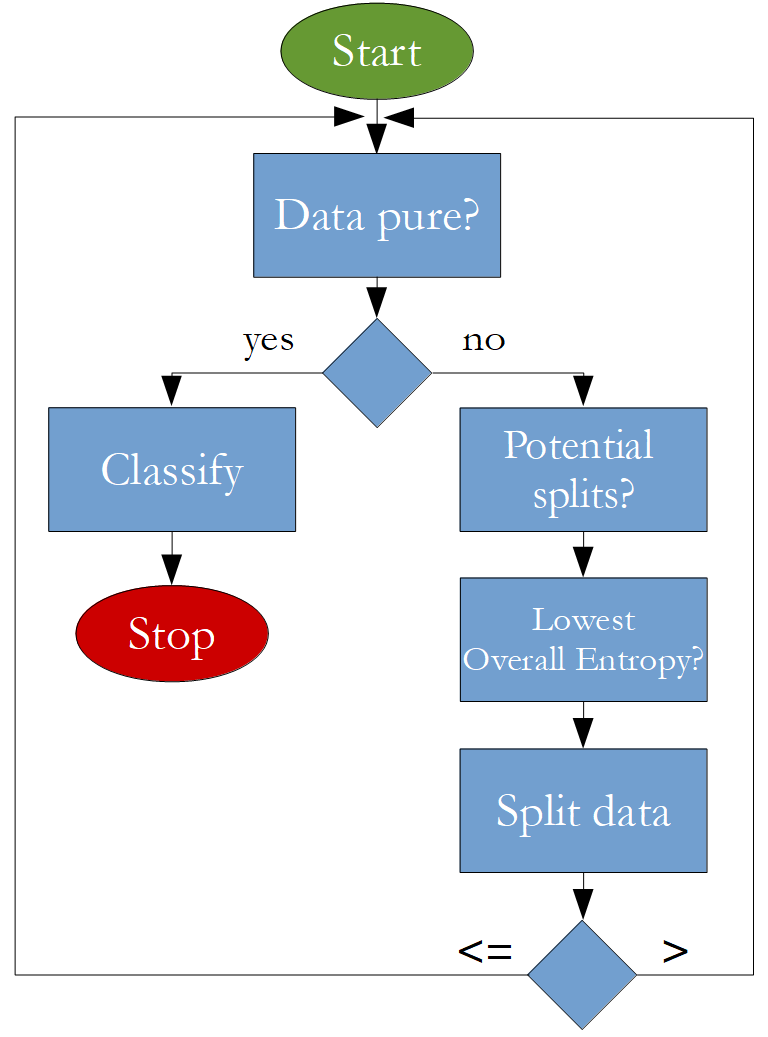
Import Statements¶
In [1]:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import random
from pprint import pprint
In [2]:
%matplotlib inline
sns.set_style("darkgrid")
Load and Prepare Data¶
Format of the data¶
- the last column of the data frame must contain the label and it must also be called "label"
- there should be no missing values in the data frame
In [3]:
df = pd.read_csv("../data/Iris.csv")
df = df.drop("Id", axis=1)
df = df.rename(columns={"species": "label"})
In [4]:
df.head()
Out[4]:
Train-Test-Split¶
In [5]:
def train_test_split(df, test_size):
if isinstance(test_size, float):
test_size = round(test_size * len(df))
indices = df.index.tolist()
test_indices = random.sample(population=indices, k=test_size)
test_df = df.loc[test_indices]
train_df = df.drop(test_indices)
return train_df, test_df
In [6]:
random.seed(0)
train_df, test_df = train_test_split(df, test_size=20)
Helper Functions¶
The helper functions operate on a NumPy 2d-array. Therefore, let’s create a variable called “data” to see what we will be working with.
In [7]:
data = train_df.values
data[:5]
Out[7]:
Data pure?¶
In [8]:
def check_purity(data):
label_column = data[:, -1]
unique_classes = np.unique(label_column)
if len(unique_classes) == 1:
return True
else:
return False
Classify¶
In [9]:
def classify_data(data):
label_column = data[:, -1]
unique_classes, counts_unique_classes = np.unique(label_column, return_counts=True)
index = counts_unique_classes.argmax()
classification = unique_classes[index]
return classification
Potential splits?¶
In [10]:
def get_potential_splits(data):
potential_splits = {}
_, n_columns = data.shape
for column_index in range(n_columns - 1): # excluding the last column which is the label
potential_splits[column_index] = []
values = data[:, column_index]
unique_values = np.unique(values)
for index in range(len(unique_values)):
if index != 0:
current_value = unique_values[index]
previous_value = unique_values[index - 1]
potential_split = (current_value + previous_value) / 2
potential_splits[column_index].append(potential_split)
return potential_splits
In [ ]: